About Us
Cell is a complicated interaction network of biomolecules and other molecules. The aim of our group is to study computationally the structures, dynamics and interactions of biomolecules and explore how biomolecules use physical laws to realize their biological functions.
Our research interests include
Biomolecule structure prediction
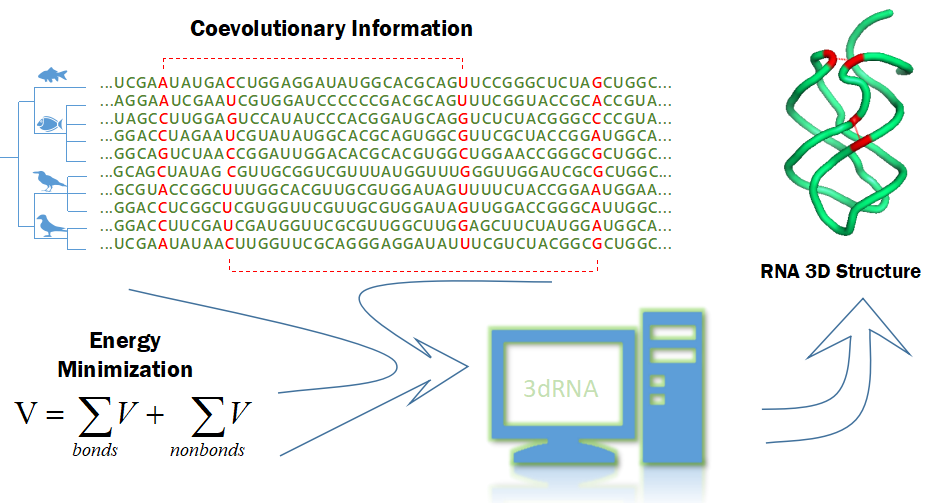
Developing computational methods of 3D structure predictions of biomolecules and their complexes. We have developed automated methods 3dRNA/DNA, 3dRPC and ASPDock for predicting 3D structures of noncoding RNAs, DNAs, RNA-protein and protein-protein complexes, respectively. We have also developed scoring functions 3dRNAscore and RPRANK for evaluating 3D RNA and RNA-protein complex structures, respectively. Currently we are trying to use physical principle and coevolutionary residue coupling and experimental contact information to improve their prediction accuracies.
 |
 |
Folding mechanisms and molecular dynamics
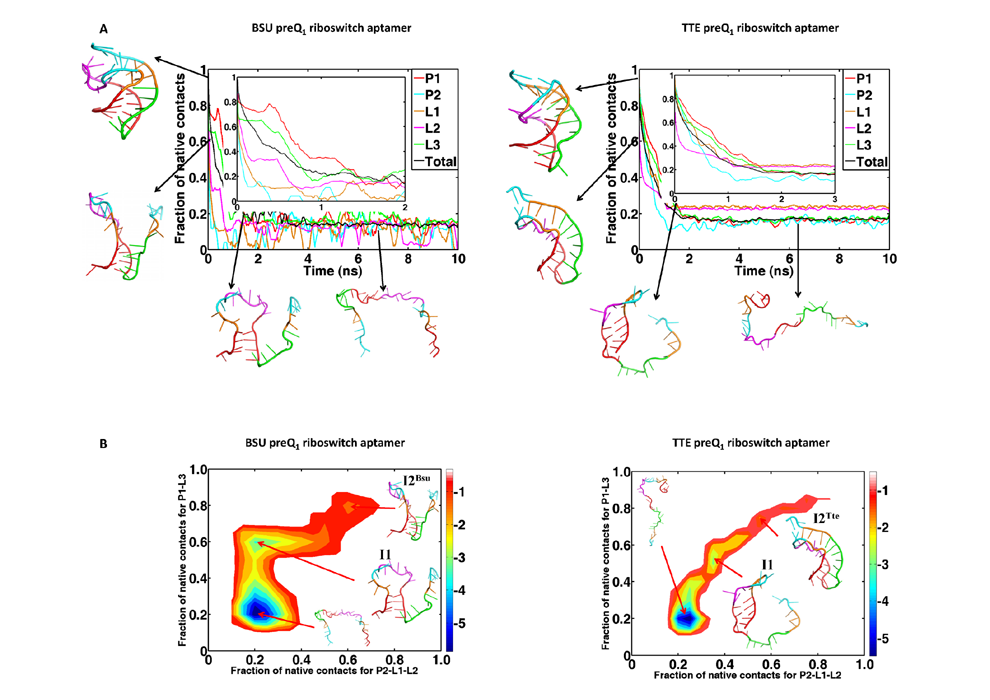
Simulating dynamics of biomolecules and their complexes and understanding their folding mechanism, dynamical behaviors and biological functions.
 |
